-
PDF
- Split View
-
Views
-
Cite
Cite
Matthew Attreed, Kristian Saied-Santiago, Hannes E Bülow, Conservation of anatomically restricted glycosaminoglycan structures in divergent nematode species, Glycobiology, Volume 26, Issue 8, August 2016, Pages 862–870, https://doi.org/10.1093/glycob/cww037
Close - Share Icon Share
Abstract
Heparan sulfates (HS) are glycosaminoglycans of the extracellular matrices and characterized by complex modification patterns owing to sulfations, epimerization, and acetylation. Distinct HS modification patterns have been shown to modulate protein–protein interactions during development in general and of the nervous system in particular. This has led to the heparan sulfate code hypothesis, which posits that specifically modified HS epitopes are distributed in a tissue and cell-specific fashion to orchestrate neural circuit formation. Whether an HS code exists in vivo, how specific or how evolutionarily conserved the anatomical distribution of an HS code may be has remained unknown. Here we conduct a systematic comparison of HS modification patterns in the nematode Caenorhabditis elegans using transgenic expression of 33 different HS-specific single chain variable fragment antibodies. We find that some HS modification patterns are widely distributed in the nervous system. In contrast, other HS modification patterns appear highly cell-specific in both non-neuronal and neuronal cells. Some patterns can be as restricted in their localization as to single neurites or synaptic connections between two neurons. This restricted anatomical localization of specific HS patterns can be evolutionarily conserved over a span of 80–100 million years in the divergent nematode species Caenorhabditis briggsae suggesting structural and, possibly functional conservation of glycosaminoglycan structures similar to proteins. These findings suggest a HS code with subcellularly localized, unique glycan identities in the nervous system.
Introduction
In metazoans, the extracellular matrices (ECMs) help to coordinate communication between cells during development and homeostasis. ECMs are complex mixtures of polymers, including proteins and glycans. For example, glycosaminoglycans of the heparan sulfate (HS) type represent a class of unbranched macromolecules (Bernfield et al. 1999) that play important roles throughout the life cycles of all metazoans (Bülow and Hobert 2006; Bishop et al. 2007; Sarrazin et al. 2011). These glycans are characterized by modifications including sulfations, epimerizations, and acetylations that are introduced during biosynthesis in the Golgi apparatus and result in extraordinary molecular complexity (Figure 1A) (Lindahl et al. 1998; Bernfield et al. 1999; Esko and Lindahl 2001). The glycan chains are invariably attached to a protein backbone to form HS proteoglycans. Because HS is not synthesized on a template, HS from biological samples displays significant molecular heterogeneity. The composition of HS changes during development and aging, and is characteristic for different tissues.
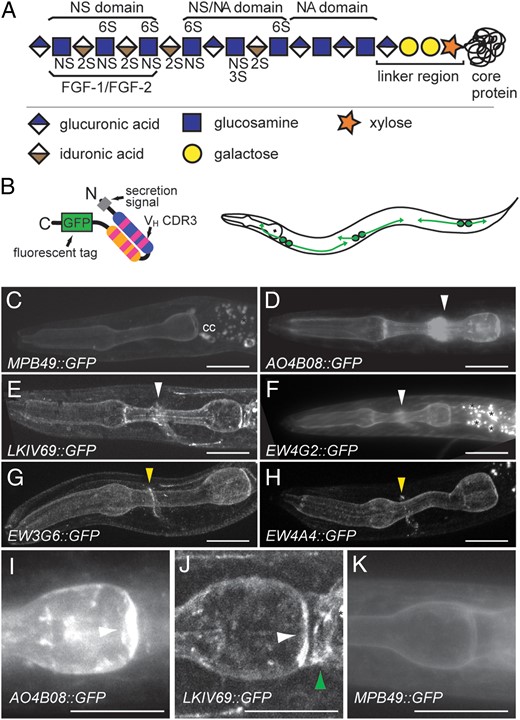
HS synthesis and anti-HS-binding patterns. (A) HS chains are attached to core proteins via an invariable tetrasaccharide linker region. Chains are non-uniformly and non-randomly modified during biosynthesis by specific enzymes in the Golgi as indicated. The resultant variably modified HS polysaccharide contains domains of varying sulfate content, including domains that can bind proteins, such as the FGF-1/FGF-2 growth factor. NS, 2S, 3S, 6S: sulfate groups in the respective positions of the sugar moieties. (B) Diagram of an scFv antibody fused to a fluorescent protein. A secretion signal targets the fusion protein for secretion (left panel). The fluorescent protein used in these experiments is the GFP(S65T) or a tandem repeat of the superfolder GFP. Schematic of a C. elegans animal transgenically secreting HS-specific scFvs into the body cavity from a set of scavenger cells termed coelomocytes (right panel). (C–H) Maximum intensity projections of optically sectioned animals transgenically expressing scFv::GFP fusions as indicated. MPB49::GFP (C) served as a negative control antibody that binds no known epitope. cc: indicate coelomocytes that stain because they reuptake unbound scFv antibody. Asterisks indicate non-specific autofluorescence. White arrowheads indicate staining of the nerve ring and yellow arrowheads denote highly restrictive expression on a single or a few neurites. Anterior is to the left and a scale bar indicates 10 µm in all panels. (I–K) Maximum intensity projections of optically sectioned animals transgenically expressing scFv::GFP fusions as indicated. MPB49::GFP (K) served as a negative control antibody that binds no known epitope. A white arrowhead denotes the pharyngeal pm8 cell, whereas a green arrow indicates the pharyngeal-intestinal valve.
HS glycans mediate protein–protein interactions in the extracellular space by way of their modification patterns (Lindahl and Li 2009; Xu and Esko 2014). Genetic loss- and gain-of-function experiments have shown that different combinations of HS modifications are required for neuronal development in both vertebrates and invertebrates (Bülow and Hobert 2006; Van Vactor et al. 2006; Poulain and Yost 2015). For example, genetic removal of enzymes that introduce HS modifications individually or in combination results in defects in retino-tectal axon projections in the optic chiasm of mice (Pratt et al. 2006; Conway, et al. 2011; Conway Price et al. 2011). Similarly, studies in worms show that different neurites rely on distinct combinations of HS modifications for various aspects of their development (Bülow and Hobert 2004). Moreover, genetic experiments show that HS modification patterns can function instructively in vivo, likely by mediating ligand–receptor interactions (Bülow et al. 2008). Collectively, these studies pointed to the existence of cell or tissue-specific HS patterns (an HS code), which guides neuronal development in metazoans by modulating protein–protein interaction (reviewed in Habuchi et al. 2004; Holt and Dickson 2005; Van Vactor et al. 2006; Poulain and Yost 2015). Whether such a code indeed exists in vivo, how specific, or how evolutionarily conserved the anatomical distribution of such an HS code may be, has remained largely unknown. Here we conduct a systematic analysis of HS modification patterns in the nematode Caenorhabditis elegans using a live imaging approach. We show that different HS modification patterns are widely distributed throughout the animals with particular diversity in the nervous system. Some modification patterns appear specifically localized to individual neurites or connections between neurites, raising the possibility of single neurite-specific glycan structures. Interestingly, some of these anatomically restricted HS modification patterns seem conserved over 80–100 million years of evolution in the nematode Caenorhabditis briggsae suggesting an important function for this structure and, providing the first example of an anatomically conserved, defined glycosaminoglycan structure.
Results
A diverse HS landscape in C. elegans
To systematically investigate HS modification patterns in live animals, we chose the small nematode Caenorhabditis elegans, with its invariant cell lineage (Sulston and Horvitz 1977; Sulston et al. 1983) and defined nervous system (White et al. 1986; Jarrell et al. 2012). We used a set of 33 HS-specific single chain variable fragment (scFv) antibodies that were previously isolated through panning of phage display libraries against different HS/heparin preparations by van Kuppevelt and colleagues (van Kuppevelt et al. 1998) (Table I; Supplementary Data). The commonly used MPB49 scFv antibody (Smits et al. 2006), which does not recognize any known epitope served as a negative control in all experiments (Figure 1C and K). While the precise HS structures recognized by the HS-specific scFvs remain largely unknown, it has been shown that the scFvs do recognize distinct epitopes in vitro (e.g. Dennissen et al. 2002). We directly visualized different HS modification patterns in live C. elegans by using a technique that involves transgenic secretion of HS-specific scFv antibody-fluorescent protein fusions into the body cavity of the animals (Figure 1B) (Attreed et al. 2012). By analyzing animals expressing each of the scFv antibodies we found about half of the HS-specific antibodies (16/33) to display labeling of the nervous system including the nerve ring (the major neuropil in the head of the worm) as well as often the ventral and dorsal nerve cords (Figure 1C–F; Table I; Supplementary data, Table SI). Staining of the nervous system was not identical with different HS-specific scFvs. Some scFv antibodies labeled the majority of neuronal tracts including the dorsal and ventral nerve cords (e.g. AO4B08, HS3A8), whereas others, such as LKIV69 or EW4G2, appeared to show more selective binding in the nerve ring (Table I; Supplementary data, Table SI). The signal was localized to neurites and essentially absent from cell somata.
Properties of HS scFv antibodies in vivo and in vitro
| scFv . | HS modifications required for bindinga . | Anatomical binding in C. elegansb . | Refs.c . | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| In vitro . | In vivo . | Nervous system . | Alimentary system . | ||||||||||||||||
| NAc . | NS . | C5 . | 2S . | 6S . | 3S . | NAc . | NS . | C5 . | 2S . | 6S . | 3S . | nr . | vnc . | dnc . | Comments . | Pharynx . | Gut . | ||
| EW4G2 | + | + | + | + | + | van de Westerlo et al. (2002) | |||||||||||||
| HS3A8 (EW3H12) | + | + | + | + | + | + | + | + | + | Gut | Attreed et al. (2012), Dennissen et al. (2002), van de Westerlo et al. (2002), Wijnhoven et al. (2008) | ||||||||
| HS4C3 | + | (+) | + | + | (+) | + | + | + | + | + | Gut | Attreed et al. (2012), Smits et al. (2006), Wijnhoven et al. (2008) | |||||||
| HS4E4 | + | + | + | + | in. | + | + | + | Dennissen et al. (2002), Wijnhoven et al. (2008) | ||||||||||
| AO4B08d | + | + | + | + | + | + | + | + | + | pm8 | Attreed et al. (2012), Dennissen (2002), Wijnhoven et al. (2008) | ||||||||
| LKIV69 | + | + | + | + | (+) | (+) | pm8, pi | Wijnhoven et al. (2008) | |||||||||||
| EW4A4 | − | + | + | +e | + | Single neuron | van de Westerlo et al. (2002), this study | ||||||||||||
| EW3G6 | (+) | + | in. | ±f | + | Single neuron | van de Westerlo et al. (2002), this study | ||||||||||||
| EV3D1 | + | + | (+) | (+) | Weak/absent | Dennissen et al. (2002) | |||||||||||||
| MPB49 | Neg. contr. | van Kuppevelt et al. (1998) | |||||||||||||||||
| scFv . | HS modifications required for bindinga . | Anatomical binding in C. elegansb . | Refs.c . | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| In vitro . | In vivo . | Nervous system . | Alimentary system . | ||||||||||||||||
| NAc . | NS . | C5 . | 2S . | 6S . | 3S . | NAc . | NS . | C5 . | 2S . | 6S . | 3S . | nr . | vnc . | dnc . | Comments . | Pharynx . | Gut . | ||
| EW4G2 | + | + | + | + | + | van de Westerlo et al. (2002) | |||||||||||||
| HS3A8 (EW3H12) | + | + | + | + | + | + | + | + | + | Gut | Attreed et al. (2012), Dennissen et al. (2002), van de Westerlo et al. (2002), Wijnhoven et al. (2008) | ||||||||
| HS4C3 | + | (+) | + | + | (+) | + | + | + | + | + | Gut | Attreed et al. (2012), Smits et al. (2006), Wijnhoven et al. (2008) | |||||||
| HS4E4 | + | + | + | + | in. | + | + | + | Dennissen et al. (2002), Wijnhoven et al. (2008) | ||||||||||
| AO4B08d | + | + | + | + | + | + | + | + | + | pm8 | Attreed et al. (2012), Dennissen (2002), Wijnhoven et al. (2008) | ||||||||
| LKIV69 | + | + | + | + | (+) | (+) | pm8, pi | Wijnhoven et al. (2008) | |||||||||||
| EW4A4 | − | + | + | +e | + | Single neuron | van de Westerlo et al. (2002), this study | ||||||||||||
| EW3G6 | (+) | + | in. | ±f | + | Single neuron | van de Westerlo et al. (2002), this study | ||||||||||||
| EV3D1 | + | + | (+) | (+) | Weak/absent | Dennissen et al. (2002) | |||||||||||||
| MPB49 | Neg. contr. | van Kuppevelt et al. (1998) | |||||||||||||||||
aHS modifications required for binding of scFvs based on competitive binding assays (in vitro) or as determined genetically (in vivo). +, required; (+) weakly required; −, not required, in., inhibitory.
bAbbreviations used: nr, nerve ring; vnc, ventral nerve cord; dnc, dorsal nerve cord; pi, pharyngeal-intestinal; pm, pharyngeal muscle; +, expressed; (+), weakly expressed.
cReferences describing original isolation and/or HS binding properties of scFvs.
dAO4B08 shows some additional staining in the M-lines of the muscle and the dense bodies.
eBoth type I and II HS 3-O-sulfotransferases are required for binding in vivo. The epitope recognized by EW4A4 appears attached to syndecan (data not shown).
fHS 3-O-sulfation introduced by type I HS 3-O-sulfotransferases acts to inhibit, whereas HS 3-O-sulfation introduced by type II HS 3-O-sulfotransferases is required for binding in C. elegans (this study). The epitope recognized by EW3G6 appears attached to both syndecan and glypican (data not shown). For a complete list of scFv fusions tested and additional information, see Supplementary data, Table SI.
Properties of HS scFv antibodies in vivo and in vitro
| scFv . | HS modifications required for bindinga . | Anatomical binding in C. elegansb . | Refs.c . | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| In vitro . | In vivo . | Nervous system . | Alimentary system . | ||||||||||||||||
| NAc . | NS . | C5 . | 2S . | 6S . | 3S . | NAc . | NS . | C5 . | 2S . | 6S . | 3S . | nr . | vnc . | dnc . | Comments . | Pharynx . | Gut . | ||
| EW4G2 | + | + | + | + | + | van de Westerlo et al. (2002) | |||||||||||||
| HS3A8 (EW3H12) | + | + | + | + | + | + | + | + | + | Gut | Attreed et al. (2012), Dennissen et al. (2002), van de Westerlo et al. (2002), Wijnhoven et al. (2008) | ||||||||
| HS4C3 | + | (+) | + | + | (+) | + | + | + | + | + | Gut | Attreed et al. (2012), Smits et al. (2006), Wijnhoven et al. (2008) | |||||||
| HS4E4 | + | + | + | + | in. | + | + | + | Dennissen et al. (2002), Wijnhoven et al. (2008) | ||||||||||
| AO4B08d | + | + | + | + | + | + | + | + | + | pm8 | Attreed et al. (2012), Dennissen (2002), Wijnhoven et al. (2008) | ||||||||
| LKIV69 | + | + | + | + | (+) | (+) | pm8, pi | Wijnhoven et al. (2008) | |||||||||||
| EW4A4 | − | + | + | +e | + | Single neuron | van de Westerlo et al. (2002), this study | ||||||||||||
| EW3G6 | (+) | + | in. | ±f | + | Single neuron | van de Westerlo et al. (2002), this study | ||||||||||||
| EV3D1 | + | + | (+) | (+) | Weak/absent | Dennissen et al. (2002) | |||||||||||||
| MPB49 | Neg. contr. | van Kuppevelt et al. (1998) | |||||||||||||||||
| scFv . | HS modifications required for bindinga . | Anatomical binding in C. elegansb . | Refs.c . | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| In vitro . | In vivo . | Nervous system . | Alimentary system . | ||||||||||||||||
| NAc . | NS . | C5 . | 2S . | 6S . | 3S . | NAc . | NS . | C5 . | 2S . | 6S . | 3S . | nr . | vnc . | dnc . | Comments . | Pharynx . | Gut . | ||
| EW4G2 | + | + | + | + | + | van de Westerlo et al. (2002) | |||||||||||||
| HS3A8 (EW3H12) | + | + | + | + | + | + | + | + | + | Gut | Attreed et al. (2012), Dennissen et al. (2002), van de Westerlo et al. (2002), Wijnhoven et al. (2008) | ||||||||
| HS4C3 | + | (+) | + | + | (+) | + | + | + | + | + | Gut | Attreed et al. (2012), Smits et al. (2006), Wijnhoven et al. (2008) | |||||||
| HS4E4 | + | + | + | + | in. | + | + | + | Dennissen et al. (2002), Wijnhoven et al. (2008) | ||||||||||
| AO4B08d | + | + | + | + | + | + | + | + | + | pm8 | Attreed et al. (2012), Dennissen (2002), Wijnhoven et al. (2008) | ||||||||
| LKIV69 | + | + | + | + | (+) | (+) | pm8, pi | Wijnhoven et al. (2008) | |||||||||||
| EW4A4 | − | + | + | +e | + | Single neuron | van de Westerlo et al. (2002), this study | ||||||||||||
| EW3G6 | (+) | + | in. | ±f | + | Single neuron | van de Westerlo et al. (2002), this study | ||||||||||||
| EV3D1 | + | + | (+) | (+) | Weak/absent | Dennissen et al. (2002) | |||||||||||||
| MPB49 | Neg. contr. | van Kuppevelt et al. (1998) | |||||||||||||||||
aHS modifications required for binding of scFvs based on competitive binding assays (in vitro) or as determined genetically (in vivo). +, required; (+) weakly required; −, not required, in., inhibitory.
bAbbreviations used: nr, nerve ring; vnc, ventral nerve cord; dnc, dorsal nerve cord; pi, pharyngeal-intestinal; pm, pharyngeal muscle; +, expressed; (+), weakly expressed.
cReferences describing original isolation and/or HS binding properties of scFvs.
dAO4B08 shows some additional staining in the M-lines of the muscle and the dense bodies.
eBoth type I and II HS 3-O-sulfotransferases are required for binding in vivo. The epitope recognized by EW4A4 appears attached to syndecan (data not shown).
fHS 3-O-sulfation introduced by type I HS 3-O-sulfotransferases acts to inhibit, whereas HS 3-O-sulfation introduced by type II HS 3-O-sulfotransferases is required for binding in C. elegans (this study). The epitope recognized by EW3G6 appears attached to both syndecan and glypican (data not shown). For a complete list of scFv fusions tested and additional information, see Supplementary data, Table SI.
Some scFv antibodies showed anatomically highly restricted staining. For example, two HS-specific scFv antibodies, EW3G6 (Attreed et al. 2012) and EW4A4, labeled what appeared to be the process(es) of only a single (or few) neurons in the nerve ring (Figure 1G and H). The observed signal localized specifically to neurites and showed an irregular punctate pattern, reminiscent of synaptic staining (Figure 1G and H). However, restricted expression of specific HS modification patterns was not limited to the nervous system. Both the AO4B08 and LKIV69 scFv antibodies detected the pharyngeal muscle cell that caps the posterior end of the pharynx (pm8), while LKIV69 also detected the pharyngeal-intestinal valve and AO4B08 labeled defined structures in body wall muscles (Figure 1I–K, Table I; Supplementary data, Table SI, not shown). Yet others labeled the gut (HS3A8, HS4C3) (Table I) or occasionally tissues of the reproductive system (not shown). The remainder of tested scFv antibodies (17/33) showed no obvious staining with the possible exception of occasional staining of the pharyngeal basement membrane, suggesting that the HS epitopes recognized by these scFvs are either not present in C. elegans or below the detection limit of the method used. Collectively, these observations showed that distinct HS modification patterns are (1) localized to different cells (including non-overlapping sets of cells) and, (2) are localized to specific subcellular compartments. The findings further suggested that subsets or, possibly single neurons are decorated with more or less unique HS modification patterns. However, this level of specificity is not limited to the nervous system, as some antibodies recognized individual cells also in the alimentary system (AO4B08, LKIV69).
Unique HS epitopes localize to the interface between the AIB and RIM interneurons
A striking example of the anatomical restriction of HS epitopes was the identification of two scFv antibodies that seemingly recognized single (or a limited number of) neurites in the head, including EW3G6 and EW4A4 (Figure 1G and H). To determine, whether the epitopes recognized by these scFv antibodies, which we term HSEW3G6 and HSEW4A4, were expressed on the same or adjacent structures in the nerve ring, we conducted a double-labeling experiment with EW3G6 and EW4A4 where the scFv antibodies were fused to a green and red fluorescent protein, respectively. We found both antibodies to display substantial but not complete colocalization in the same region of the nerve ring (Figure 2A). To determine whether the HSEW3G6 and HSEW4A4 epitopes are structurally similar or distinct, we crossed animals transgenically expressing the respective scFvs with loss-of-function mutations in HS genes. We found that both EW3G6 and EW4E4 required HS 2-O-sulfation for binding (Figure 2B and C, Table I). Moreover, HS 6-O-sulfation and HS 3-O-sulfation introduced by the type II HS 3-O-sulfotransferase hst-3.2 were required for binding of EW4A4 to its epitope(s) (Figure 2C, Table I). In contrast, HS 6-O-sulfation and HS 3-O-sulfation introduced by the type I HS 3-O-sulfotransferase hst-3.1 was inhibitory for binding of EW3G6 to its cognate epitope(s) (Figure 2B; Table I). These experiments establish that the epitopes recognized in vivo by EW4A4 and EW3G6 constitute HS. Moreover, these observations imply that the HSEW3G6 and HSEW4A4 epitopes are molecularly distinct entities that decorate closely juxtaposed cellular structures, although we cannot rule out indirect effects of removing individual modifying enzyme on the stability, trafficking, or localization of HS epitopes.
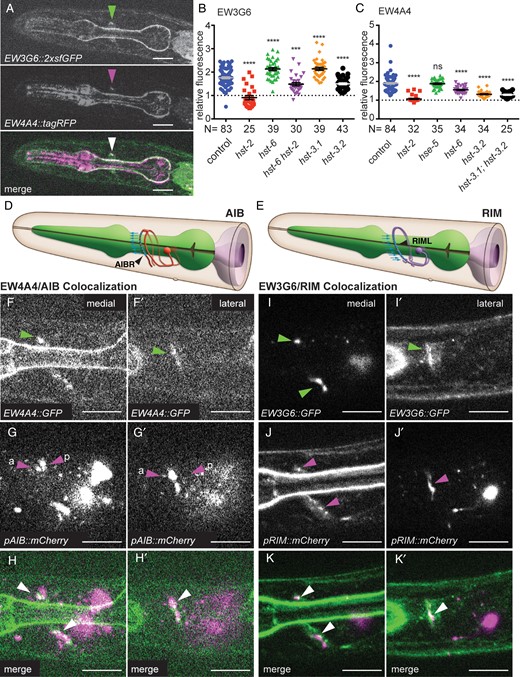
EW3G6::2xsfGFP and EW4A4::2xsfGFP bind distinct epitopes in proximity to the synaptic connection between AIB and RIM interneurons in the C. elegans nerve ring. (A) Fluorescent optical sections of the same focal plane of animals expressing both EW3G6::2xsfGFP (green channel, upper panel) and EW4A4::tagRFP (middle panel) with labeling of epitopes indicated by a green or magenta arrowhead, respectively. A merged image (lower panel) shows partial colocalization (white arrowhead). Anterior is to the left. Scale bars in all panels indicate 10 µm. (B and C) Quantification of relative fluorescence of transgenic strains expressing the EW3G6::2xsfGFP (dzEx1031) and EW4A4::2xsfGFP (dzEx1042) fusions in mutant backgrounds as indicated: hst-2, HS 2-O-sulfotransferase; hst-6, HS 6-O-sulfotransferase; hst-3.1, HS 3-O-sulfotransferase (type I); hst-3.2, HS 3-O-sulfotransferase (type II); hse-5, HS C5-epimerase. Statistical significance compared with the respective wild-type controls was calculated using the unpaired t-test with Welch's correction and indicated as ****, P <0.0001; ***, P <0.001, ns, not significant. Errors bars denote the standard error of the mean. (D and E) Schematics of AIB interneurons (left panel) and RIM motor/interneurons (right panel) with synaptic output and input between both, respectively, indicated by blue arrows. Modified after www.wormatlas.org, accessed 28 March 2016. (F–H′) Fluorescent optical sections of a medial/sagittal (F–H) and a lateral/parasagittal plane (F′–H′) of animals transgenically expressing the EW4A4::2xsfGFP and mCherry as a cytoplasmic marker in the bilateral pair of AIBL/R neurons under control of the Pinx-1 promoter (Altun et al. 2009). Images of the green (E and E′) and red (F and F′) channels with a merged image (H and H′) are shown. Green and red arrowheads point to labeling of the respective processes and indicate the more anterior descending (a), and posterior ascending (p) neurites of AIB, respectively. White arrowheads point to colocalization. Anterior is to the left, ventral down. The reciprocal experiment between EW4A4 and RIML/R is shown in Supplementary data, Figure S2C–E′. Fluorescent optical sections of a medial/sagittal (I–K) and a lateral/parasagittal plane (I′–K′) of animals transgenically expressing the EW3G6::2xsfGFP and mCherry as a cytoplasmic marker in RIML/R under control of the Pgcy-13 promoter (Ortiz et al. 2006). Similar results were obtained with the Ptdc-1 promoter, which is also expressed in RIM (Alkema et al. 2005). Images of the green (I and I′) and red (J and J′) channels with a merged image (K and K′) are shown. Green, red and white arrowheads point to labeling of the respective processes and colocalization. The reciprocal experiment between EW3G6 and AIBL/R is shown in Supplementary data, Figure S2F–H′.
Aided by knowledge of the anatomical structure of the C. elegans nervous system (White et al. 1986) and the availability of cell-specific promoters, we set out to determine which cell or cells express the HSEW3G6 and HSEW4A4 epitopes. We systematically labeled defined cellular processes in the nerve ring with cytoplasmic fluorescent markers in one color in animals that secreted the EW4A4 (or EW3G6) scFv fusions with a differently colored fluorescent protein (Supplementary data, Table SII). For example, we found both the HSEW3G6 and HSEW4A4 signals always positioned anterior and somewhat more medial to the neurites of AIY interneurons in the nerve ring (Supplementary data, Figure S1). On the other hand, we found the HSEW3G6 and HSEW4A4 signals consistently more dorsal and slightly more posterior than the neurites of RIA interneurons (Supplementary data, Figure S1). By systematically conducting similar experiments with different promoters driving cytoplasmic fluorescent markers, and the scFv antibody-GFP fusions we determined the approximate positions of the HSEW3G6 and HSEW4A4 epitopes by triangulation (Supplementary data, Figure S1). We found that both HSEW3G6 and HSEW4A4 staining appeared in very close proximity to the neurites of RIML(left)/R(right) and AIBL(left)/R(right) interneurons (Figure 2D–K′; Supplementary data, Figure S2), consistent with the earlier observation that both antibodies display some colocalization. The neurites of the bilateral pair of AIB neurons (AIBL(left)/R(right)) normally project from the cell soma towards the ventral cord, where they run anteriorly before turning dorsally as part of the nerve ring around the pharynx. At the dorsal midline, the neurites both turn anteriorly for a short distance to then descend on the opposite site of the animal where they form synapses with RIML(left)/R(right) (Figure 2D, Supplementary data, Figure S2) (White et al. 1986). In contrast, the neurites of the pair of RIM neurons travel up in the nerve ring—after a brief dip into the ventral cord—and directly descend on the opposite site (Figure 2E) (White et al. 1986). The ascending neurites of the RIMs receive synaptic input from the descending neurites of the contralateral AIB, i.e. the RIML neurite receives input from AIBR and RIMR from AIBL (Figure 2D and E; Supplementary data, Figure S2G). In colocalization experiments, it appeared as if the EW3G6 signal displayed partial overlap with RIM neurites while being closely opposed to the anterior descending neurites of AIB neurons. Conversely, the EW4A4 signal displayed partial overlap with the anterior descending neurites of AIB neuron while being closely apposed to the neurites of RIM neurons (Figure 2D–K′; Supplementary data, Figure S2). In an attempt to delineate the origin of the HSEW3G6 and HSEW4A4 epitopes, we investigated the expression pattern of hst-2 and hst-6 reporter transgenes (Bülow and Hobert 2004). We found that both pairs of RIM and AIB neurons expressed a hst-6::GFP reporter, whereas a hst-2::GFP reporter was expressed only in RIM, but not in AIB neurons (Figure 3). Given that hst-2 activity is required for binding of the EW3G6 and EW4A4 scFv antibodies to their respective epitopes (Figure 2B and C), these findings imply that the HSEW3G6 and HSEW4A4 epitopes are produced by RIM interneurons. In conclusion, we find that two molecularly distinct HS epitopes, which likely originate from RIM neurons, are localized to a region within the nerve ring that is in close apposition with the synaptic connections between AIB and RIM neurons. Together with the survey of HS modification patterns these observations indicate that specific HS epitopes are distributed in C. elegans with unexpectedly high anatomical specificity to possibly defined connections between neurons.
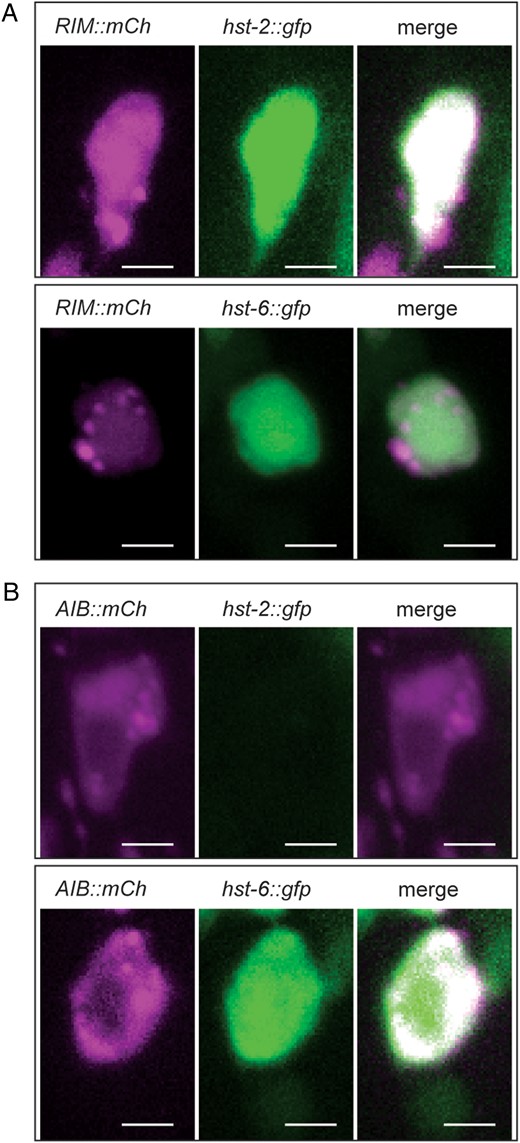
RIM and AIB neurons express a hst-6::GFP reporter but only RIM neurons express a hst-2::GFP reporter. (A) Fluorescent optical sections of the same focal plane of cell bodies in animals expressing a RIM cytoplasmic marker (Pgcy-13::mCherry) (red channel, left panels) and a hst-2::GFP or hst-6::GFP cytoplasmic reporters, respectively (middle panels). Merged images are in right panels. The RIM neurons appear to be the only pair of neurons with strong expression of the hst-2::GFP reporter in the head region whereas the hst-6::GFP reporter displays widespread expression in many additional head neurons at different levels. Three independent transgenic lines gave comparable results for each combination. Scale bars indicate 2 µm in all panels. (B) Fluorescent optical sections of the same focal plane of cell bodies in animals expressing an AIB cytoplasmic marker (Pinx-1::mCherry) (red channel, left panels) and hst-2::GFP or hst-6::GFP cytoplasmic reporters, respectively (middle panels). Merged images are in right panels. Three independent transgenic lines gave comparable results for each combination.
Anatomically restricted HS epitopes are evolutionarily conserved
The conservation of protein structure, and of specific amino acid residues within a protein during evolution is a sign of functional significance, and has been extensively studied since similar amino acid sequences for insulin from different species were first reported (Brown et al. 1955). In contrast, much less is known about the conservation of non-template-based glycan structures. Generally, glycans are remarkable for their structural diversity throughout evolution rather than for structural conservation (Gagneux and Varki 1999; Varki 2011). Nonetheless, the enzymatic machinery involved in the biosynthesis of glycans is highly conserved from invertebrates to vertebrates, including all enzymes involved in the biosynthesis of HS (Häcker et al. 2005; Bülow and Hobert 2006). Consistent with the conservation of the biosynthetic machinery, HS is detected already in planarians and widely distributed throughout the animal kingdom (Yamada et al. 2011). Moreover, the fact that the scFvs, which we have been using were originally isolated against HS and heparan from vertebrate sources, clearly shows that similar HS epitopes exist in vertebrates and invertebrates. Whether the anatomical localization of HS epitopes can be conserved, has not been determined.
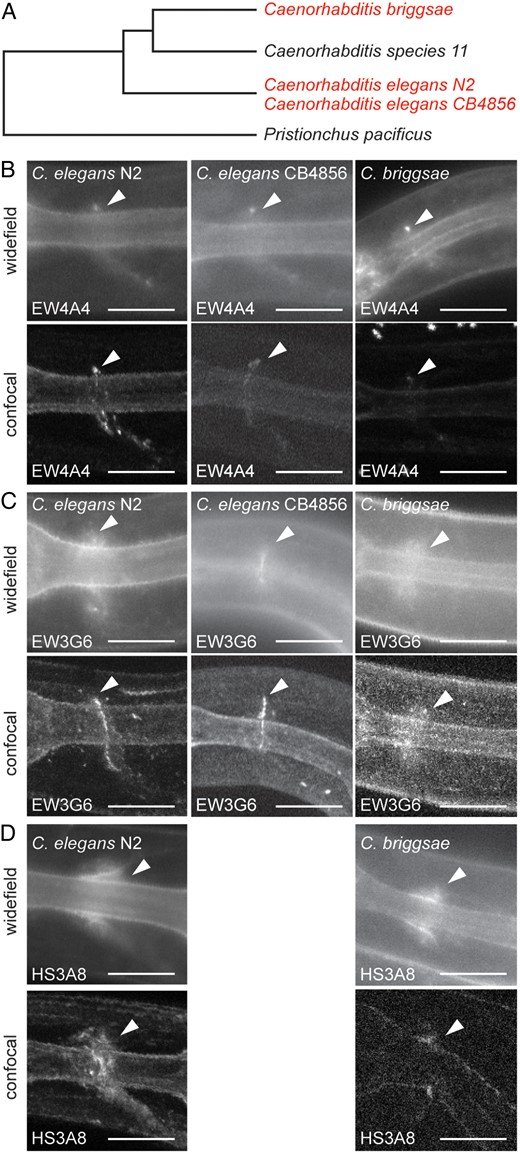
Our finding of spatially highly restricted and molecularly distinct HS epitopes afforded us the opportunity to test whether such specific localization is evolutionarily conserved and to what degree. We used two additional nematode strains, a C. elegans isolate from Hawaii (CB4856) which shows partial incompatibility with C. elegans (N2) and may be at the cusp of speciation (Seidel et al. 2008), and the related species Caenorhabditis briggsae (AF16) which diverged ∼80–100 million years ago from C. elegans, a time frame similar to the divergence of mice and humans (Figure 4A) (Stein et al. 2003). We compared staining patterns in transgenic animals of all three strains/species expressing the cell-specific antibodies EW3G6 and EW4A4 as well as a more broadly staining antibody, HS3A8. We found the labeling patterns of the EW4A4 scFv antibody to a single neurite(s) indistinguishable between C. elegans (N2), C. elegans (CB4856), and C. briggsae (Figure 4B), suggesting that the highly restricted anatomical distribution of the rare HSEW4A4 epitope is conserved between C. elegans and C. briggsae (Figure 4B). In contrast, the labeling patterns of the EW3G6 antibody to a single neurite was conserved in the Hawaiian isolate of C. elegans but not in C. briggsae where we found a broader distribution. Thus, the anatomical localization of HSEW3G6 was not completely conserved, although this or a closely related epitope was still present and localized to the nervous system (Figure 4C). Finally, we tested the distribution of a more common epitope in C. elegans, HSHS3A8, and found a similar distribution in C. briggsae as in C. elegans (Figure 4D). In conclusion, we have detected some evolutionary change in subcellular localization patterns of HS epitopes (e.g. HSEW3G6), but also a surprising conservation of an anatomically restricted HS epitope (HSEW4A4) in C. briggsae, which is as distant from C. elegans as mice from men.
Evolutionary conservation of anatomically restricted HS patterns. (A) Dendrogram indicating the evolutionary relationship between different nematode strains of the genus Caenorhabditis with Pristionchus as outgroup. Species used in experiments are shown in red. N2 and CB4856 are C. elegans wild-type isolates from Bristol (UK) and Hawaii (USA), respectively, that are partially incompatible and may be at the cusp of speciation (Seidel et al. 2008). (B–D) Fluorescent widefield and maximum intensity projections of optically sectioned animals expressing EW4A4::2xsfGFP (B), EW3G6::2xsfGFP (C) or HS3A8::2xsfGFP (D) scFv fusions in Caenorhabditis sp. as indicated. White arrowheads indicate apparently conserved labeling of a single neurite in the head of the animals in all species whereas empty arrowheads point to obviously divergent labeling patterns. Anterior is to the left in all images and ventral down. Scale bars indicate 10 µm. This figure is available in black and white in print and in color at Glycobiology online.
Discussion
Specificity of scFv antibodies
We present here the first attempt of a systematic survey of the ‘HS landscape’ in a whole animal using the majority of available HS-specific single chain variable antibodies (Supplementary data, Table SI). Our studies show overlapping and distinct localization of different HS epitopes. The high number of antibodies that recognize HS epitopes in the nervous system is consistent with the prior observation that the majority of HS in C. elegans is localized to the nervous system (Minniti et al. 2004). Yet, HS is not exclusive to the nervous system and we also find anatomically restricted localization of HS epitopes to parts of the pharynx and muscle. It was unexpected to find two scFv antibodies with distinct binding specificity to bind in such close proximity and partial overlap to, possibly, a specific synaptic connection. One possibility is that both scFvs recognize partially overlapping epitopes, although we consider this possibility less likely, given that they behave differently genetically (Figure 2B and C) and they display non-overlapping localization in C. briggsae. Alternatively, but not mutually exclusive, resolution of our method may be insufficient to separate adjacent HS epitopes on the same HS chain or HS epitopes carried by different HS core proteins in close proximity.
The heparan sulfate code
Analyses of the heparan sulfate code have primarily focused on neuronal guidance (Poulain and Yost 2015). However, early work suggested that HS and HS proteoglycans are important for establishment of functional synapses, in particular with regard to the neuromuscular junction (Poulain and Yost 2015). Recent work in humans, mice, and drosophila further suggested that HS is important for the establishment of correct synaptic circuits and cognitive function. Mutations in human EXT1, a gene involved in HS biosynthesis, have been associated with autism (Li et al. 2002) and, mice defective for Ext1 display behavioral deficits reminiscent of autism spectrum disorders (Irie et al. 2012). In drosophila, knockdown of the HS 6-O-sulftaransferase resulted in synaptic defects (Dani et al. 2012). Interestingly, the staining patterns we observe with different scFvs appear often punctate and excluded from the cell soma, reminiscent of synaptic staining. Consistent with this interpretation, HS-specific staining has previously been shown in at least one case to colocalize with presynaptic markers in C. elegans (Attreed and Bülow 2015). The genes encoding enzymes required for introducing modifications in the HSEW3G6 and HSEW4A4 epitopes (including hst-2, hst-6 and hst-3.2) are not obviously required for patterning of RIM and AIB axonal projections (data not shown). Thus, we speculate that specific HS modification patterns are not only important for guiding neurites during development, but may also play a role in the establishment, function or maintenance of specific synaptic connections. Since neuronal connectivity and, in many cases, the function of certain connections is known in C. elegans, this model may represent a unique opportunity to distinguish between these possibilities in future experiments.
Conservation of glycosaminoglycan structures
The experiments presented here establish that defined HS epitopes are (a) localized with unexpected anatomical specificity in the nervous system and (b) that this localization appears conserved, at least in some cases, over 80–100 million years of evolutionary time. Glycans of different types are present on all cellular surfaces and are essential for life (Varki 2011). In contrast to proteins where synthesis is template driven, glycans are synthesized by non-template driven processes. Moreover, whereas proteins are often conserved during evolution and allow for the deduction of time scales when a last common ancestor was alive, glycans are much more diverse and generally less conserved (Varki 2011). This makes sense in light of many of their biological functions, which include for instance the recognition of self (or foreign). Such diversity may go as far as intraspecies variability of cell surface antigens, such as for instance the ones that distinguish blood serotypes (reviewed in Gagneux and Varki 1999), although, interestingly, this intraspecies diversity in and of itself appears conserved in apes (Martinko et al. 1993). Nonetheless, examples do exist where glycan structures appear conserved, such as the N-glycan structure of different mammalian plasma fibrinogens or, pituitary glycoproteins (reviewed in Gagneux and Varki 1999). Some indirect evidence suggests that the heparin–antithrombin interaction may be conserved from amphibians to humans (Jordan 1983), implying that the heparin structure is conserved as well. However, the example presented here is to our knowledge the first instance where anatomical localization of specific glycosaminoglycan structures has been documented on an evolutionary time scale.
Materials and methods
Strains and transgenesis
Worms were cultured as previously described (Brenner 1974). The following strains and alleles were used: N2 Bristol C. elegans wild-type, CB4856 Hawaiian C. elegans polymorphic strain, and AF16 Gujarat C. briggsae wild-type. For transgenesis in C. briggsae we used DNA constructs with C. elegans regulatory sequences. Several lines of evidence suggest that they are expressed as predicted in C. briggsae. First, we do see expression of fluorescence in the C. briggsae coelomocytes, which are readily identifiable suggesting that the coelomocyte-specific Punc-122 promoter operates as expected. Second, 4/4 recently tested C. elegans promoters showed comparable expression in corresponding cells in C. briggsae (Zhao et al. 2010). Third, development and cell lineage between both species is remarkably similar in general and with regard to AIBL/R and RIML/R neurons and the nervous system in particular (Zhao et al. 2008).
To create transgenic animals, single chain variable fragment (scFv) antibody constructs were injected with pRF4 (rol-6(su1006)) as a dominant injection marker at 25 and 100 ng/µl, respectively. Alternatively, Pttx-3::mCherry or other cell-specific markers were used with the wild-type copy of dpy-20 for injection into dpy-20(e1282) mutant animals at 25 ng/µl and up to 100 ng/µl, respectively. For EW3G6 and EW4A4 lines and all transgenic lines in C. briggsae, the complex array method was used (Evans 2006). Plasmids were first linearized, and then added so that each linearized plasmid totaled 5 ng/µl in the injected solution and digested N2 DNA was used as carrier DNA. Several transgenic lines giving comparable results were obtained for each construct. All mutants and integrated transgenes were backcrossed at least four times prior to analysis. For a detailed strain list, see Supplementary Material.
Molecular biology
All scFv antibody fusion constructs were prepared as previously described (Attreed et al. 2012). Variable heavy chain cDNAs for the respective scFv antibodies were established by site-directed mutagenesis of the CDR3 in existing variable heavy chain cDNAs (Attreed et al. 2012) or de novo synthesis. Cell-specific cytoplasmic markers were produced using the PCR fusion technique (Hobert 2002). Primer sequences are available upon request. For details on molecular biology, see Supplementary data, Table S2.
Fluorometric and microscopic analyses
Microscopic images were acquired with an AxioCam MRm camera mounted on an AxioImager Z1 compound microscope using Axiovision 4.8 Zeiss software. Some worms were optically sectioned using the Zeiss Apotome on an AxioImager Z1. The Apotome uses structured illumination to acquire optical sections comparable with a scanning laser confocal microscope (Weigel et al. 2009). Optical sections were processed as 12-bit grayscale images using Zeiss Axiovision software 4.8 (if obtained with the Apotome) to produce maximum intensity projections. For preparation of figures, images were exported from Axiovision 4.8 by applying a 5% best-fit gamma curve and assembled into figures using Adobe Photoshop and Adobe Illustrator.
Quantitative epifluorescence
For quantitative epifluorescence microscopy, animals were synchronized by hypochlorite bleaching and plated on regular nematode plates with OP50 Escherichia coli as a food source. Approximately 24 h later, worms were washed off in M9, anaesthetized with 4 mM levamisole and 0.3 M 2,3-butanedione monoxime, mounted on a 5% agarose pad and analyzed on an AxioImagerZ1 compound microscope (Zeiss). All images used for quantification were taken with an exposure time of 100 ms at ×400 magnification and fluorometrically analyzed using Zeiss AxioVision software version 4.8 as follows. Relative pixel intensity in a region of ‘specific fluorescence’ was measured and divided by the relative pixel intensity measured in a region of ‘background fluorescence’ located near the posterior pharyngeal bulb to yield relative fluorescence. Statistical significance was calculated using the two-tailed Student's t-test with Welch's correction using the PRISM® software package (v.6) from Graphpad.
Authors’ contributions
M.A. and H.E.B. designed the experiments. M.A. and K.S.S. conducted the experiments and analyzed the data with H.E.B. M.A. and H.E.B. wrote the paper with editorial input from K.S.S.
Funding
This work was supported in part by grants from The National Institutes of Health (T32 GM007491 to M.A. and K.S.S. and, F31 NS076243 to M.A.; RC1 GM090825 and R01 GM01313 to H.E.B.; P30 HD071593 and P30 CA013330 to Albert Einstein College of Medicine). H.E.B. is an Alfred P. Sloan and an Irma T. Hirschl/Monique Weill-Caulier Research Fellow.
Conflict of interest statement
None declared.
Abbreviations
ECMs, extracellular matrices; HS, heparan sulfate; scFv, single chain variable fragment.
Acknowledgements
We thank S. Emmons, R. Townley and members of the Bülow lab for comments on the manuscript and helpful discussions and O. Hobert for reagents.
References